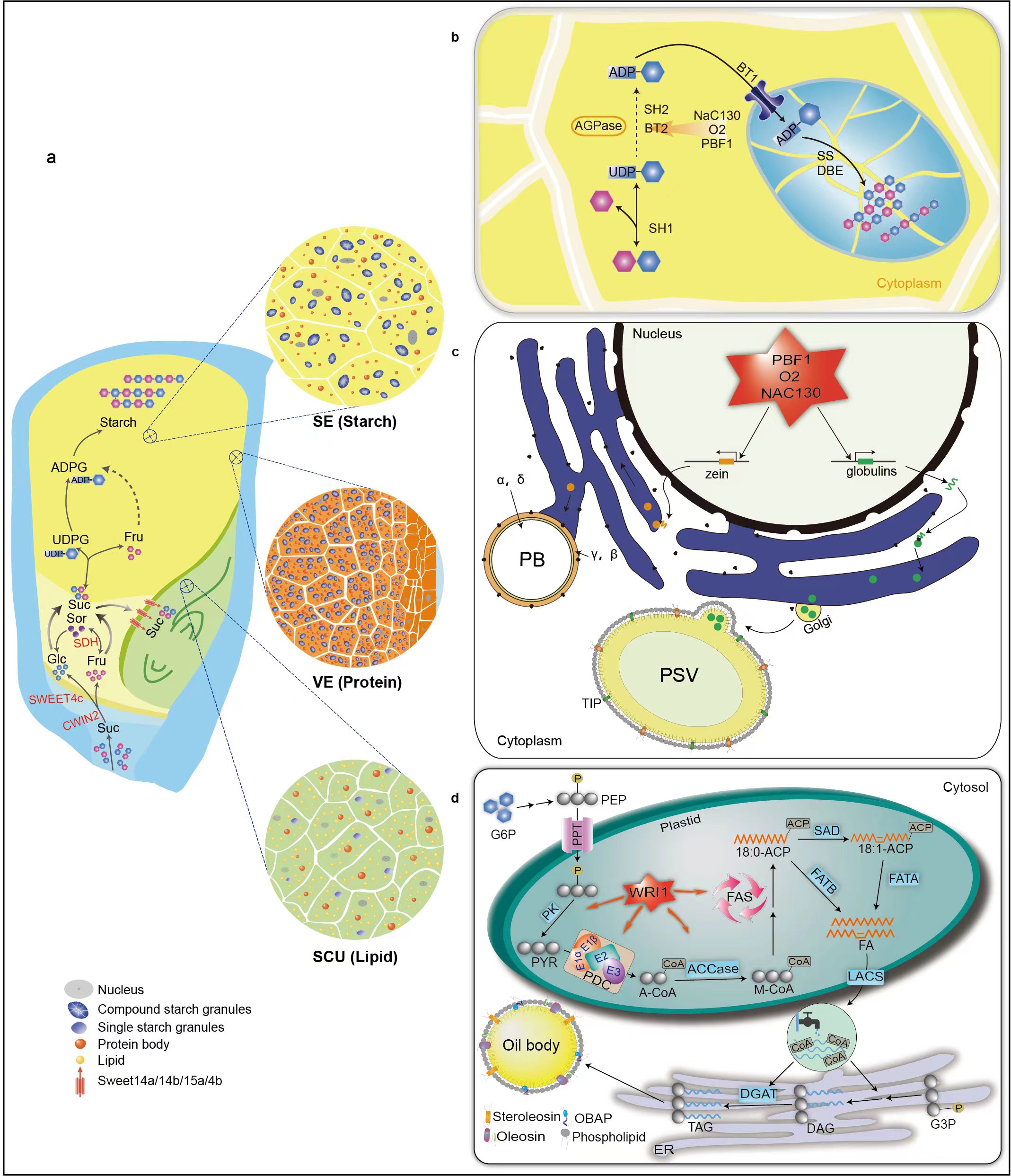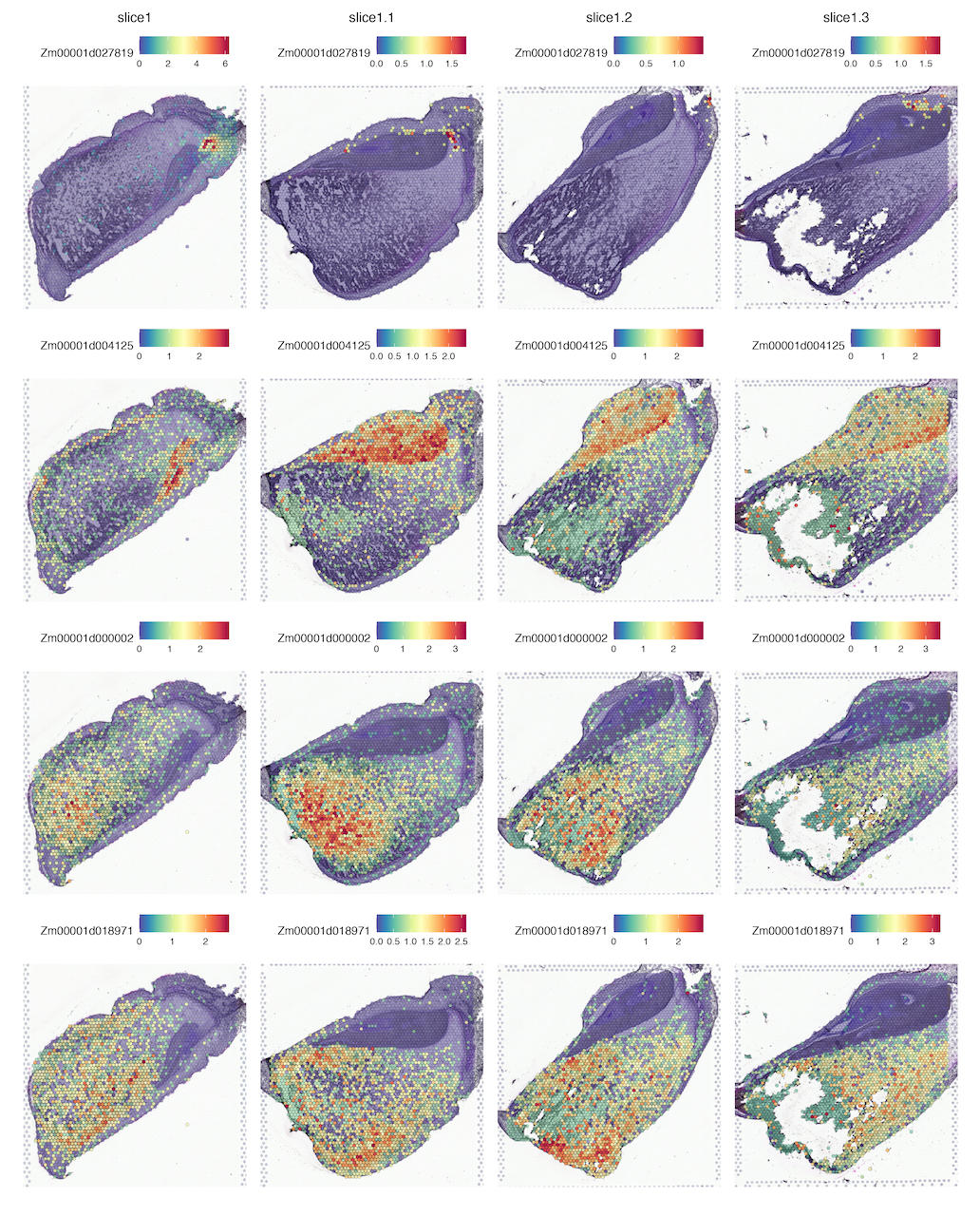


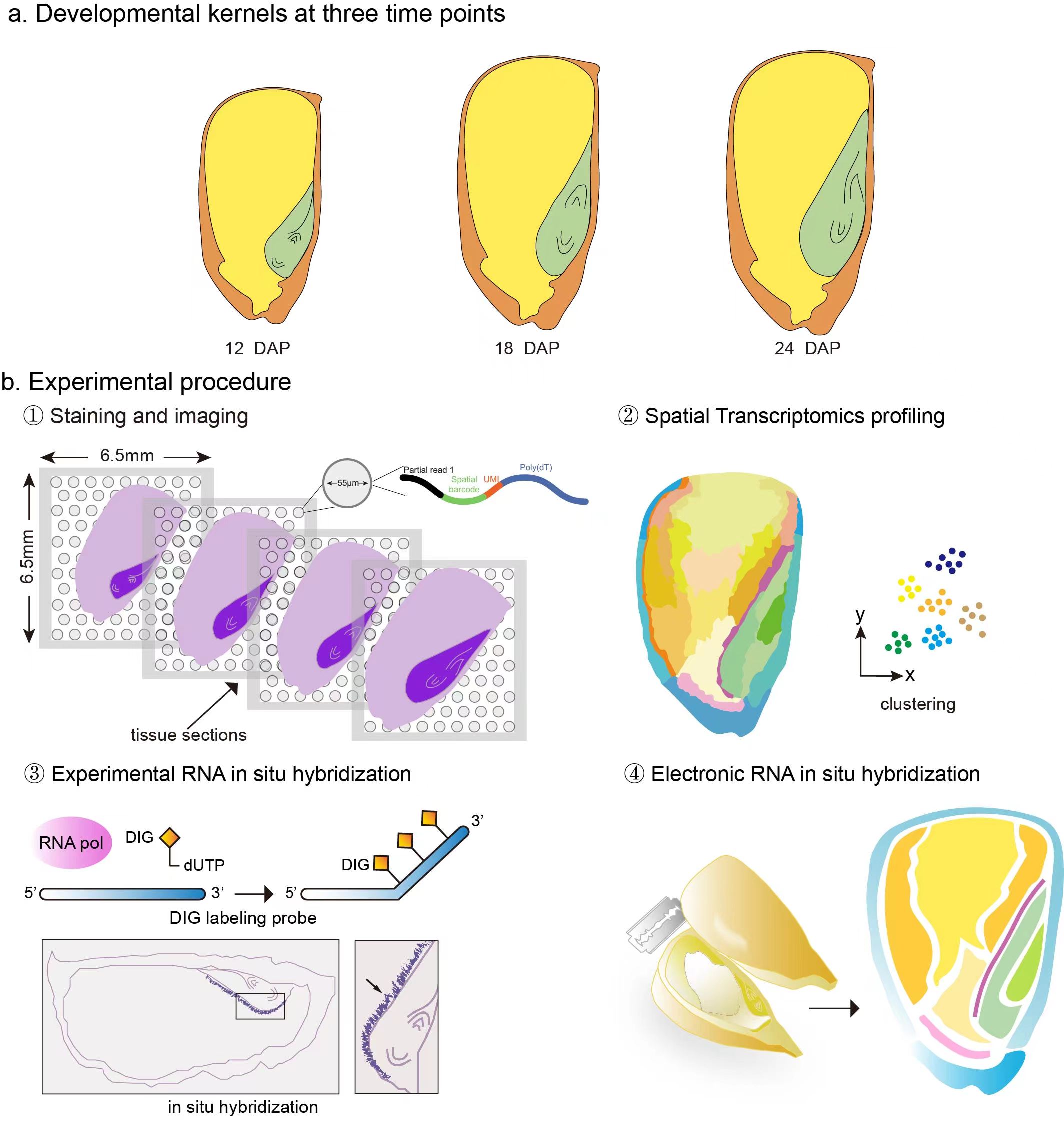
Maize kernel is a complex biological system comprising three genetically distinct tissues nested one inside another (maternal tissue, embryo and endosperm) that are responsible for protein, starch and lipid accumulation. These complex events are the result of intricate and well-timed spatially functional programs in the specialized tissue domains of kernels, whereas little is understood of the function of biological systems without access to spatially gene expression measurement.
Here, we will provide you with convenient access to visualized spatial transcription results on the page of the Electronic in situ images. You can obtain the expression levels and spatiotemporal information of your interested genes by submitting its gene identifier. The data used in this application are publicly available on the page of the About. Notics: The dataset used in the website has been standardized !
Tips: If you want to know more about gene expression information, please collaborate with us.
In the near future, we will conduct more maize kernel spatial transcriptome research, attempting to decipher the gene regulatory networks during maize kernel development. We sincerely look forward to your feedback on the construction of our website's database.
The raw sequencing data were deposited with the accession number of PRJCA015607 (https://ngdc.cncb.ac.cn/bioproject/browse/PRJCA015607). Details of the spatial transcriptomic analysis pipeline and R coding scripts can be found at Github: https://github.com/wwq413/SpatialTranscriptomics.